About My Lab's Research
Leaders in Conservation Genetics
-
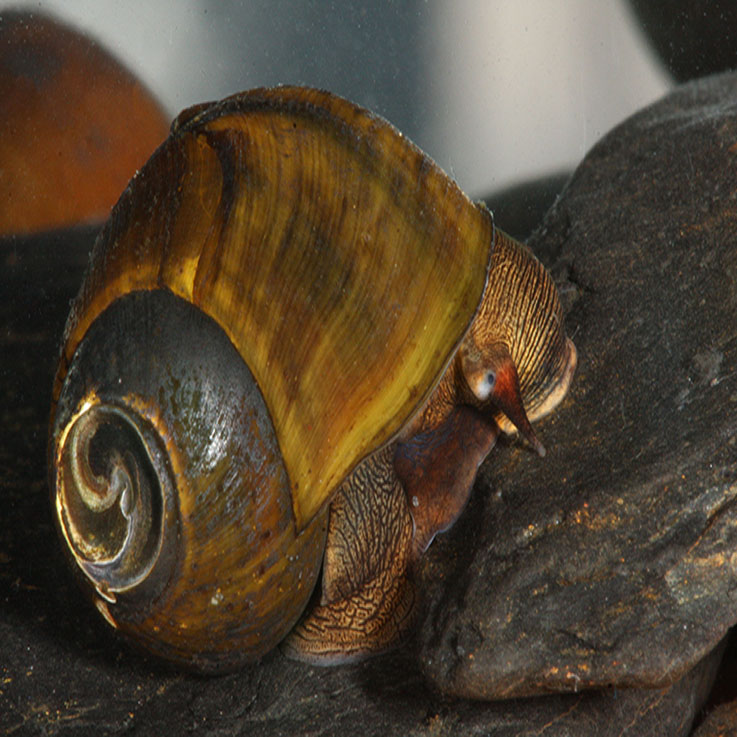

Freshwater Mollusks
Current ProjectsFreshwater mollusks are the lab's primary study organisms. One reason we study mollusks is to enhance conservation efforts. The health of many freshwater ecosystems, and therefore much of society, rely on freshwater snails and mussels. Unfortunately, over 75% of freshwater mussels and snails are at risk of extinction, and the greatest barrier to effective conservation is a lack of knowledge. We use phylogenetics and conservation genomics to better understand freshwater mollusks.
-
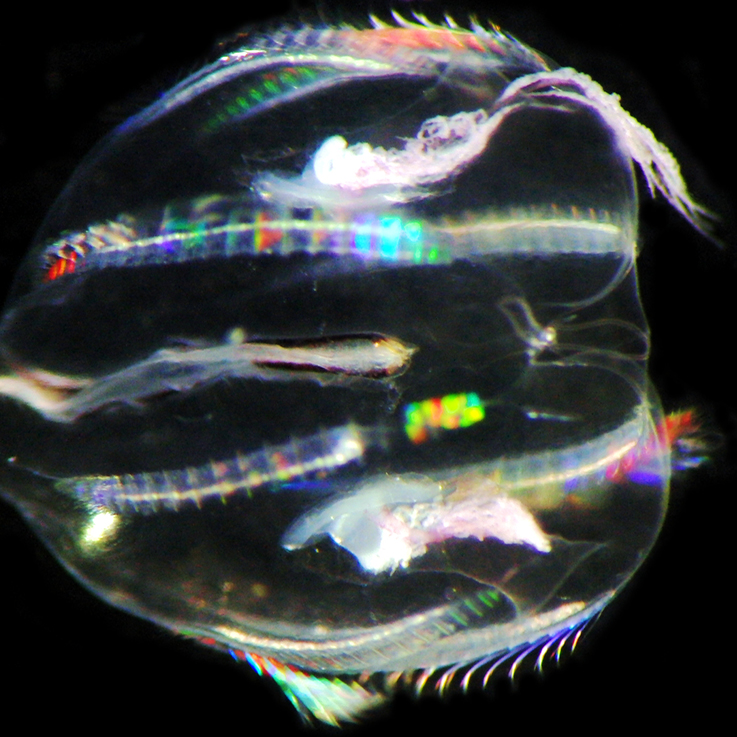

Phylogenetics and Comparative Biology
Current ProjectsMy lab seeks to understand the patterns and processes that have contributed to animal diversification. Our primary study organisms are invertebrates. We have a number of ongoing projects on phylogenomics of freshwater mollusks and conservation genomics of animals ranging from freshwater snails to sturgeon. We are also interested in evolution of gene families, particulary those involved with invertebrate adaptation to freshwater environments.
-
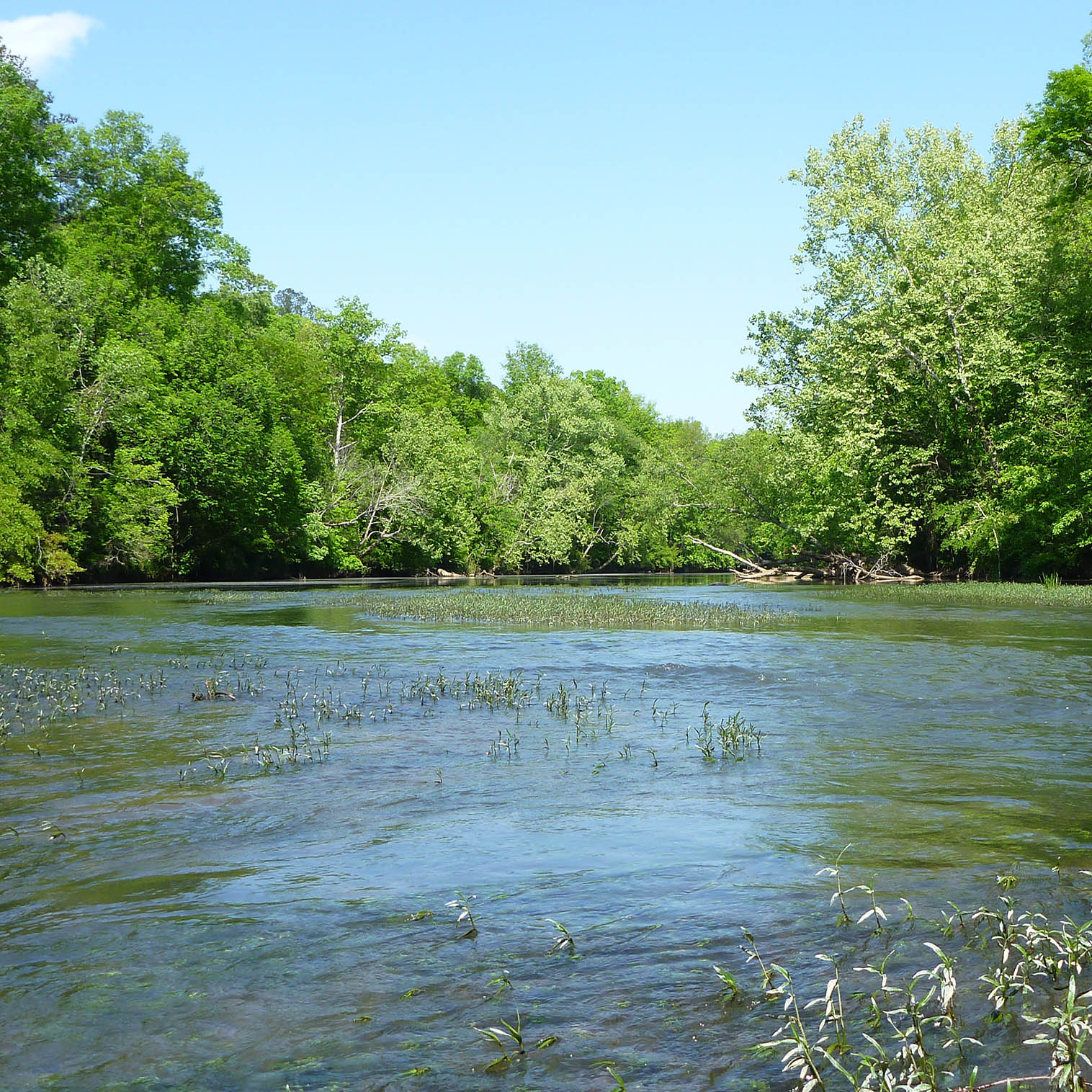

Conservation Genetics
Current ProjectsA consequence of species decline is often a decline in population-level genetic diversity. Our research aims to understand how conservation efforts can maintain or increase genetic diversity of managed populations and species. We also seek to better understand the molecular ecology of threatened and endangered species. For this research, our lab collaborates with landowners and hatcheries. We also spend time in the field sampling at-risk, threatened, and endangered species, with a focus on rivers in the southeastern United States.
-

Environmental DNA
Current ProjectsMy lab has recently expanded so we can perform cutting-edge environmental DNA (eDNA) work following best practices. eDNA research can be a cost-effective approach for oranismal surveys. We use DNA sampled from the environment to detect invasive and rare species when traditional sampling is difficult or inefficient.
-
Bioinformatics and Phylogenetic Inference
Modern genetics research requires a considerable amount of computational biology. Part of my lab's research includes designing new bioinformatics pipelines for evolutionary biology research. We also study the performance of different phylogenetic methods in an effort to determine which models and methods can provide the most accurate inferences about organismal relationships.












